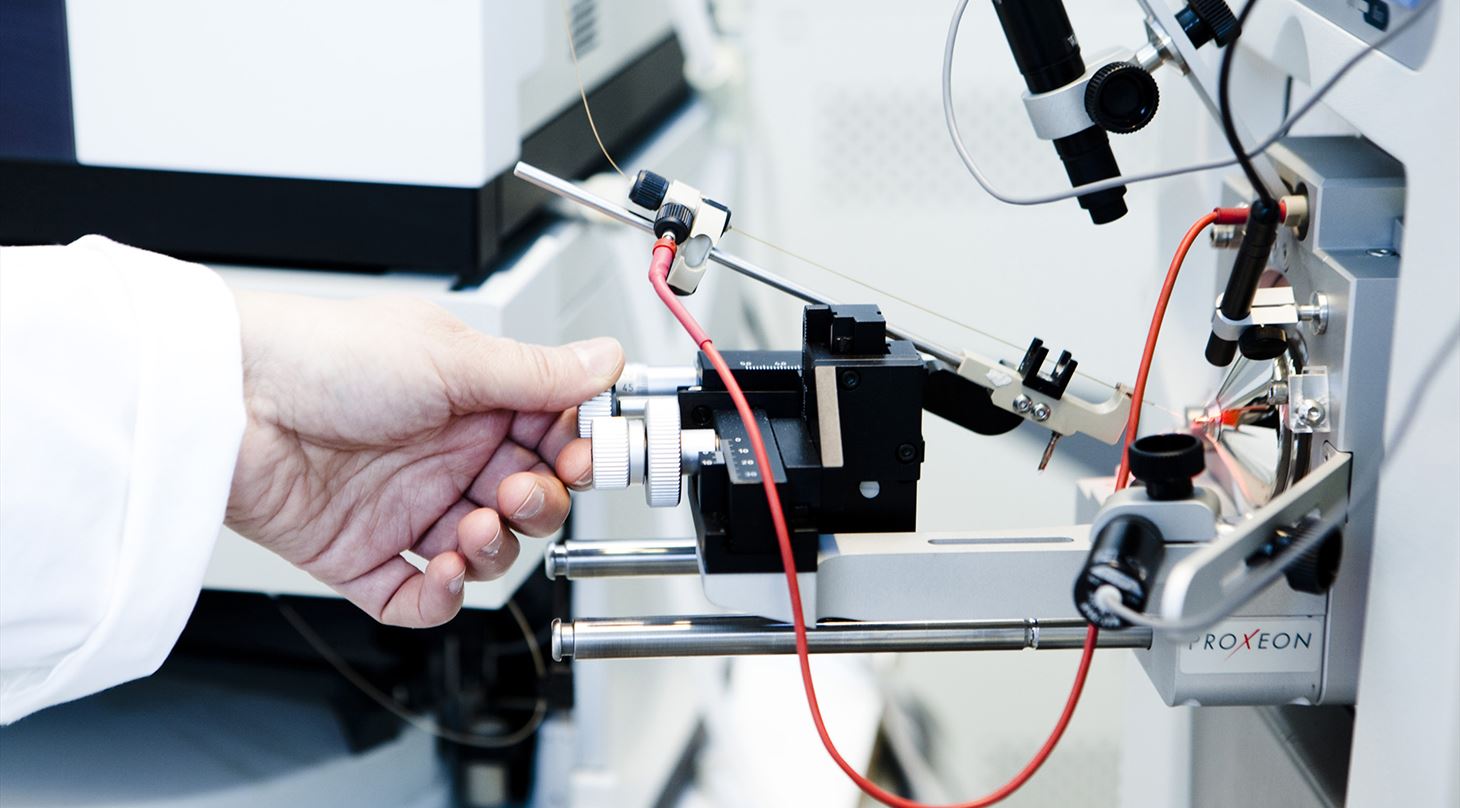

Protein characterisation, proteomics and mass spectrometry (MS)
Danish Technological Institute has extensive experience in carrying out protein characterisation and proteome analyses, including the use of advanced mass spectrometry. Our experience spans a wide range of research and development projects that Danish Technological Institute carries out in collaboration with universities and companies.
How can Danish Technological Institute help you?
We offer a range of services from robust and classical protein analyses that can provide information on the complexity and size of proteins to advanced mass spectrometry-based analyses.
 Mass spectrometry analysis
Mass spectrometry analysis
Within protein-based mass spectrometry, our analyses cover protein identification, peptide mapping, quantification and modifications, among other things.
 Protein identification
Protein identification
We can identify proteins in your product or unknown protein from your development / research. Using mass spectrometry, we can identify and quantify proteins in for example tissue samples or gels. We can identify individual proteins from gels, for example, or whole proteomes from tissue samples and cell lysates among others.
 Peptide mapping
Peptide mapping
We can confirm your protein sequence or determine the sequence of unknown proteins.
 Quantification
Quantification
We offer several for quantifying peptides and proteins in your samples. We can do total protein analysis (total nitrogen), amino acid analysis and mass spectrometry-based quantification. Using mass spectrometry, we can make absolute or relative quantification both with and without stable isotope labelling. Therefore, mass spectrometry can often be used to validate other quantification methods (e.g. ELISA) or to identify biomarkers.
 Modifications
Modifications
We can assist with the identification of modifications in full proteomes and / or the characterisation of individual proteins and their post-translational modifications (PTMs) such as phosphorylation, N-termini and disulphides, for example. We can also help find your modification and its specificity.
 Protein extraction, separation and purification
Protein extraction, separation and purification
It can often be an advantage to separate complex samples before an advanced protein analysis. We can help with this separation using SDS-PAGE and HPLC. We can also purify recombinant proteins or proteins from natural sources. We have extensive experience in the development and optimisation of protein extraction from complex samples. We can therefore help you develop an optimised extraction procedure that can increase the success of your final results. For example, we can help with the purification of glycopeptides and phosphopeptides.
 Protein expression
Protein expression
Sometimes a protein of interest is needed in certain amounts. Wi input from you, we construct protein sequences to fit your downstream requirements, produce it in a relevant expression organism, and purify it for further analyses and applications. Inclusion of specific tags, such as myc or 6xHis may simplify downstream RnD processes. Typical amounts range from a few micrograms to several grams, which will be validated in terms of activity and/or structure.
Examples of mass spectrometry usage
|
